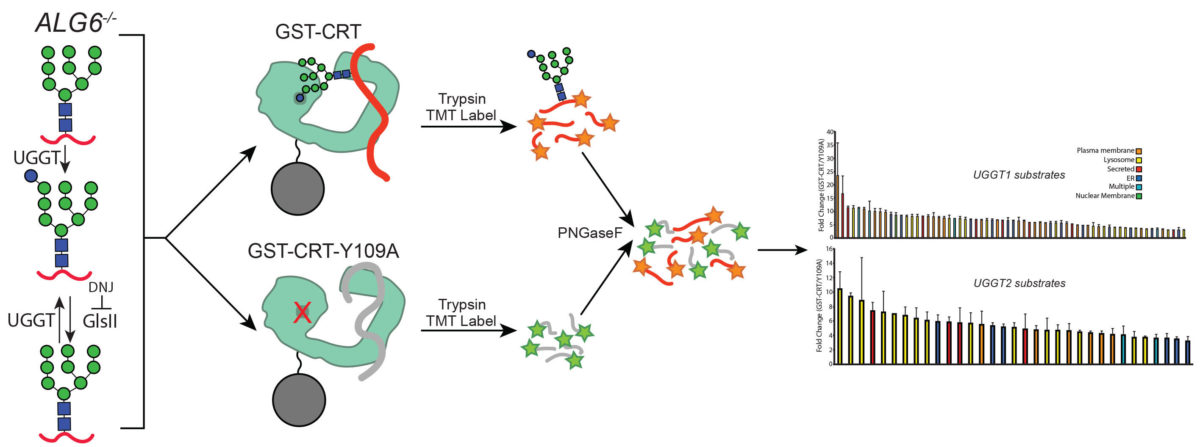
UGGT1 and UGGT2 are key quality control factors that determine the fate of glycoproteins in the early mammalian secretory pathway. These two paralogues direct persistent molecular chaperone binding in the endoplasmic reticulum that helps with protein maturation and sorting. Persistent chaperone binding of terminally misfolded clients can target proteins for degradation by the proteasome, as well as lysosomal proteases. In our study, a quantitative glycoproteomics strategy was developed to identify cellular clients of UGGT1 and UGGT2. Interestingly, UGGT1 was found to preferentially recognize large membrane glycoproteins, while UGGT2 favored the modification of smaller soluble lysosomal proteins. This study opens the door to a detailed understanding of the recognition process for these important quality control factors and identifies targets whose folding trajectories may be altered in disease states. This work, recently published in eLife, blends chemical and cell biological approaches and was a collaborative effort by three CBI students in the lab, spearheaded by lead author Ben Adams. Ben recently started a postdoctoral position at Harvard Medical School. – Dan Hebert
“Our work identified native substrates of an essential protein quality control process and explored it’s role in glycoproteostasis.” – Ben Adams
“Novel substrates of the glucosyltransferases UGGT1 and UGGT2 have been identified and compared to an in silico N-glycoproteome” – Nathan Canniff
“Through a quantitative mass spectrometry approach we have determined endogenous substrates for UDP-glucose:glycoprotein glucosyltransferases (UGGT) 1 & 2, showing both are functional and preferentially target different substrates requiring folding assistance from calnexin and calreticulin” – Kevin Guay
Quantitative glycoproteomics reveals cellular substrate selectivity of the ER protein quality control sensors UGGT1 and UGGT2.Adams BM, Canniff NP, Guay KP, Larsen ISB, Hebert DN. Elife. 2020 Dec 15;9:e63997. doi: 10.7554/eLife.63997.PMID: 33320095. Free PMC article